Swiss BioRef
Project consortium (active contribution): Alexander Leichtle (Insel), Jivko Stoyanov (Swiss Paraplegic Research), Martin Hersberger (Kispi Zürich), Nicolas Rosat (CHUV), Christos Nakas (Insel).
Supporters: Christian Lovis (UniGE/HUG), Nicolas Vuilleumier (HUG), Nazanin Sédille-Mostafaie (Insel), François Mach (HUG), Jerome Dauvillier (SIB), Baris Gencer (HUG), Idris Guessous (HUG), Thomas McKee (HUG).
Project contributors and endorsement:
- SIB and DCC: Support with setting up the consortium agreement and project governance.
- DCC: Support with setting up the Swiss BioRef ontology, including new concepts and terminologies.
- BioMedIT / SIS: Security assessment and test deployment of MedCo, developer consulting, deployment of validation tool including reverse proxy-solution for online access, support with converting external terminologies to RDF.
- Tune Insight: Integration of Swiss BioRef-related enhancements of MedCo into TI4Health codebase, efforts towards deploying TI4Health in the BioMedIT framework.
- Unitectra: Support with project governance.
Main achievements
Reference intervals for laboratory test results are in standard use across many medical disciplines, allowing physicians to pinpoint potentially pathological test results with relative ease. The process of inferring cohort-specific reference intervals, however, is often ignored due to the high costs and cumbersome effort associated with such a task. Determining reference intervals based on data collected during daily clinical routine using fully automated computational resources may help to lower the affiliated costs and personalize the reference intervals to the respective cohort population.
During the SwissBioRef project, we have developed a multi-center computational framework where specialized web applications estimate and assess patient group-specific reference intervals based on clinical routine data from four major Swiss clinical centers:
- University Hospital Bern (Inselspital)
- University Hospital Lausanne (CHUV)
- University Children’s Hospital Zurich (KiSpi)
- Swiss Paraplegic Research (including Swiss Paraplegic Center in Nottwil and University Hospital Balgrist in Zürich)
We have established a common legal and interoperability framework for our clinical partners to share their data either by transferring it via the secure BioMedIT network to a central database or by providing their data in a decentralized manner (see Figure 1). The latter option is employing MedCo (and its successor TI4Health), a secure and encrypted data-sharing system, allowing each data provider to comply with the restrictions laid out by their cantonal ethics committees.
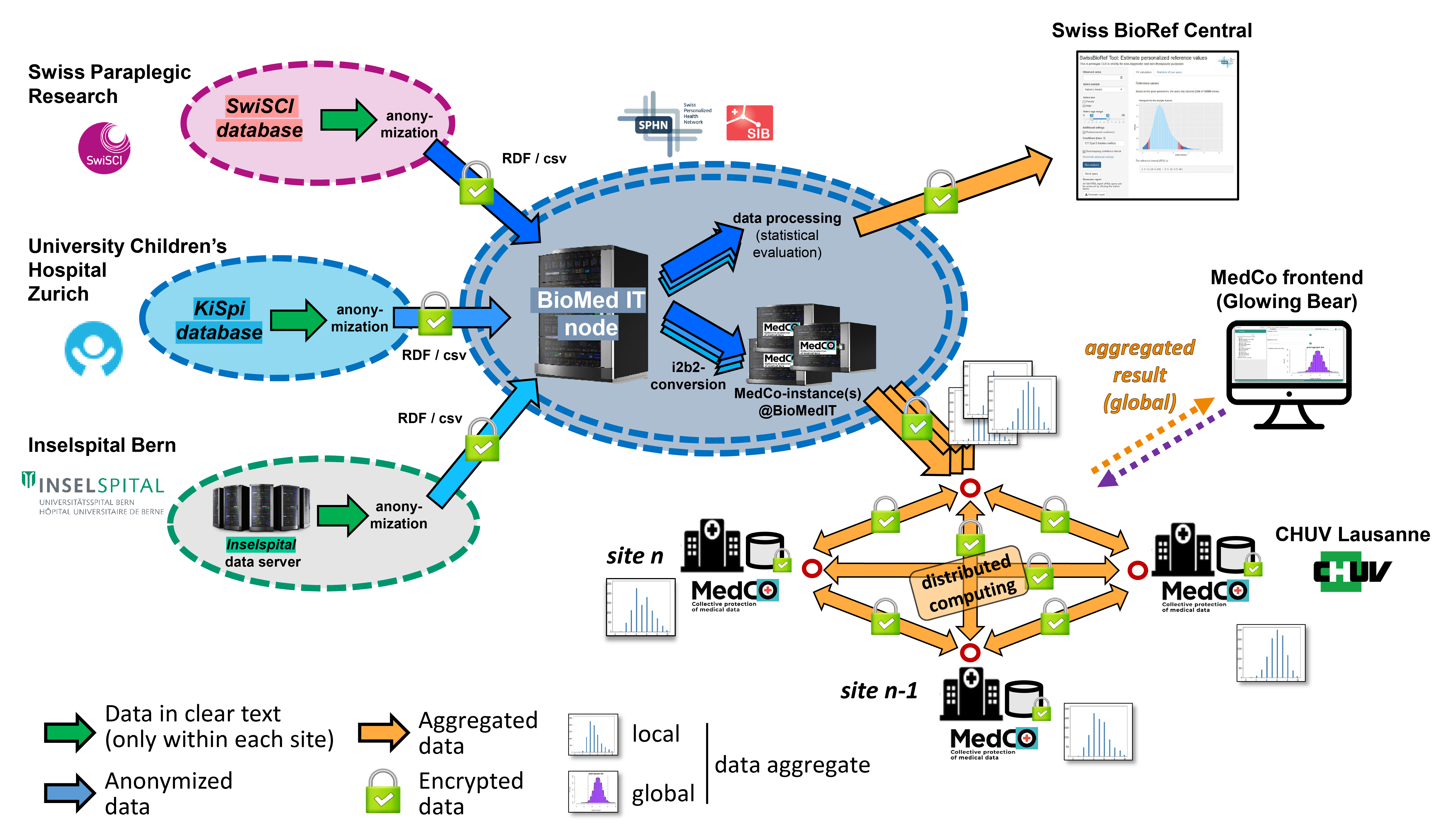
Figure 1: Schematic representation of the Swiss BioRef infrastructure
The deployed web applications allow intuitive and interactive data stratification by patient factors (like age, administrative sex and personal medical history) and laboratory analysis factors (unique identifiers for device, analyzer and test kit used for the analysis). The applications are accessible for Swiss physicians and researchers by SWITCH-edu ID.
Reusable datasets and infrastructures
1. Standardized dataset
During the Swiss BioRef project a large interoperable multi-cohort dataset has been compiled from four major hospitals in Switzerland (University Hospital Bern, University Hospital Lausanne, Swiss Paraplegic Research, and University Children’s Hospital Zurich).
The data have been harmonized in accordance with the interoperability standards of the DCC, e.g., using LOINCs as identifiers of the laboratory analyses or units according to UCUM. Inselspital has been supporting KiSpi, Swiss Paraplegic Research, and CHUV with the introduction or assignment of LOINCs for the analyses of interest where they had still been missing.
The dataset entails approx. 9 million individual measurements from approx. 250’000 patients and more than 40 analyses from clinical chemistry, hematology, point-of-care-testing and coagulation. The cohort data of Inselspital, KiSpi, and Swiss Paraplegic Research has been combined centrally within the BioMedIT framework. CHUV is keeping their cohort data separate to be made available via the decentralized access route using TI4Health.
The analyses included in the dataset are available in Table 1.
Several patient factors (age, administrative gender and ICD-10-GM-coded diagnoses) have been extracted along with the laboratory measurements. Additional analytical factors enrich the data set with metainformation concerning the analysis method and help to overcome the lack or sparsity of information LOINCs offer with respect to the applied method. This includes type identifiers or unique identifiers of analyzers or test kits used in the clinical laboratories from the Global Medical Device Nomenclature (GMDN) and Global Unique Device Identifier Database (GUDID). In a major effort, Inselspital has supplied LOINC-UDI mapping tables to the other consortium members after identifying their laboratory setups with the help of the local laboratory specialists.
Available resources
Contact persons:
Alexander Leichtle, Harald Witte
Use of the data set is currently restricted to member institutions of the Swiss BioRef consortium.
Access criteria are determined by the BioRef Executive Board.
Data contributions of the individual data providers:
| Data provider | measurements | patients | cases | unique LOINCs | time span of measurements |
| Inselspital | 5’969’733 | 186’265 | 323’600 | 39 | 06/2014 - 05/2022 |
| KiSpi | 454’215 | 17’179 | 28’393 | 37 | 04/2014 - 05/2022 |
| Swiss Paraplegic Research | 25'634 | 615 | 615 | 36 | up to 03 / 2022 |
| CHUV | 2’541’930 | 45'591 | 97'054 | 48 | 01/2020 - 03/2022 |
| Total | 8’965’878 | 249’650 | 449’662 |
2. Validation tool for estimation of reference values (“Swiss BioRef Central”), incl. advanced statistics modules
The validation tool serves as a development and validation tool for the Swiss BioRef team to test the performance and accuracy of four statistical methods on inferring precise reference intervals from multi-cohort resources. The application was developed at Inselspital for use on a single cohort data set and has since been adapted to host data from multiple cohorts. The modifications for Swiss BioRef included several functional and user experience enhancements. The back-end was modified to allow data loading and data filtering based on user input from a multiple cohort data set. For the inference of reference intervals, two direct methods (IFCC/CLSI approved) and two indirect methods (using newer data mining techniques) were implemented, with the most appropriate method being chosen automatically. The frontend was completely revised to have a user-friendly query selection process with intuitive button functionalities (such a run analysis, reset query, generate report and logging out). The result of the reference interval estimation is displayed in a prominent text box (with confidence intervals) and graphically supported with a histogram and can be exported in form of an ISO-17025 compliant HTML report (reporting functionality). Under advanced settings, users are able to change the statistical method used or define laboratory parameters for fine-grained filtering, such as the medical device or test kit ID (using unique product identifiers from the Global Unique Device Identification Database (GUDID)) and the medical device or the test kit ID type (using identifiers from the Global Medical Device Nomenclature (GMDN)). In order to simplify deployment, the application has been fully dockerized with TLS/SSL encryption on all frontend-backend traffic. All modifications have been done based on the open source framework of R, R Shiny and open source ressources. The Swiss BioRef Central application is deployed on LeoMed (SIS ETHZ) and can be accessed after registration via rshiny-swissbioref.leomed.ethz.ch/.
Available resources
Contact persons:
Alexander Leichtle, Tobias Blatter
Documentation Swiss BioRef Central (access and user manual): https://docs.google.com/document/d/1hQudR8r1PpQhM5mCOIzA1Zr_iLwy2PGWX-E_vD0_uho/edit
Validation tool (2FA-access after registration): rshiny-swissbioref.leomed.ethz.ch/
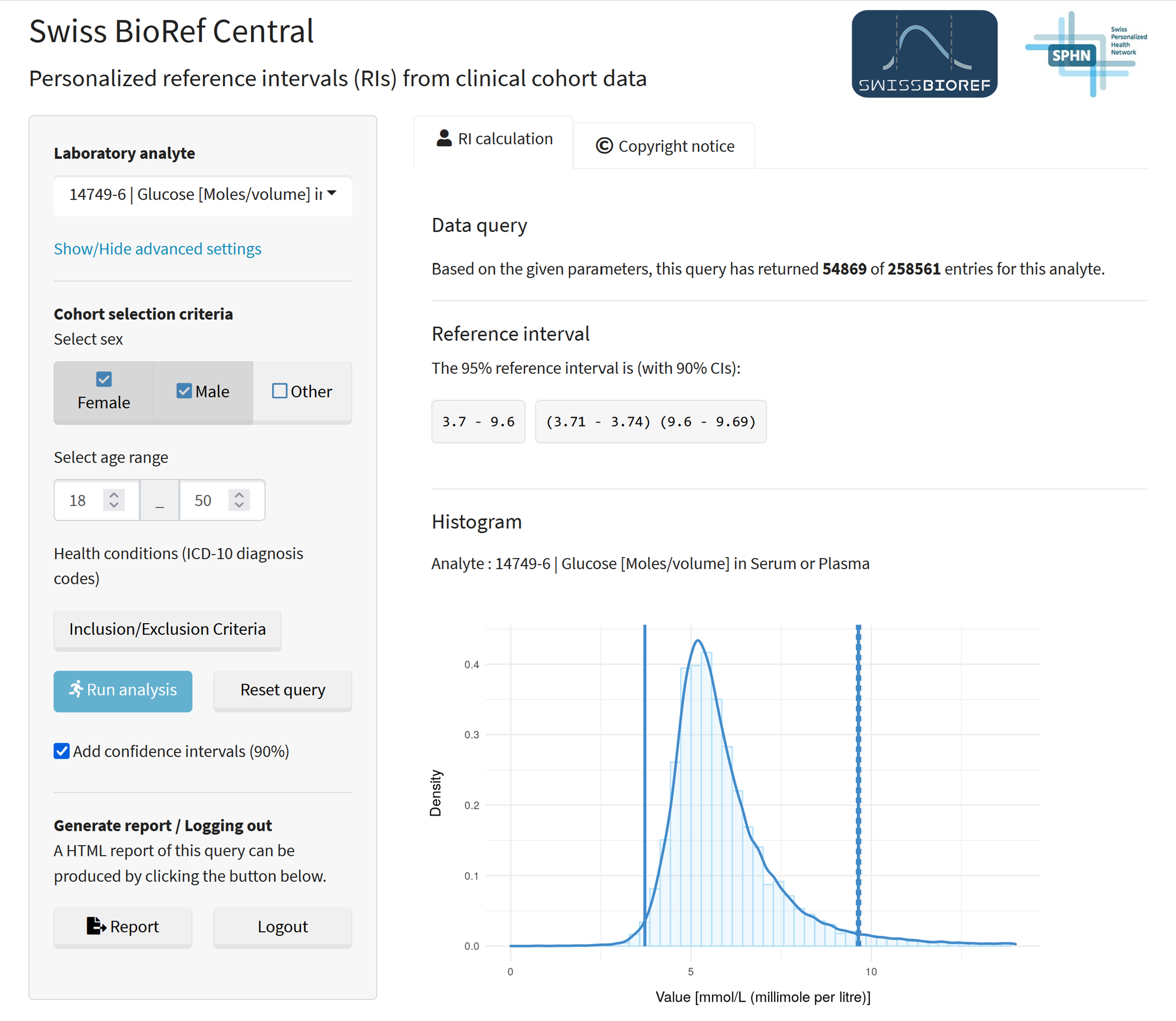
Figure 2: Swiss BioRef Central - User Interface after running a patient query, showing the calculated reference intervals.
3. Swiss BioRef project ontology, incl. new external terminologies
The ontology for Swiss BioRef is based on the SPHN ontology (release 2021-2). It has been set up at Inselspital, in close collaboration with the DCC and RDF experts.
New concepts and terminologies have been introduced in the Swiss BioRef ontology:
- Age concept (time elapsed since birth of the individual)
- LabAnalyzer concept (laboratory analyzer used to assess medical laboratory samples)
- new external terminologies (GUDID, GMDN, EMDN)
The Age concept allows an assignment of a patient’s age to a test result. This works independent of sensitive data (e.g., date of birth) facilitating the inclusion of age information in data requests.
The LabAnalyzer concept enriches analysis methods with product and type identifiers (compare the L4CHLAB report1). This additional metadata shall help to overcome the lack or sparsity of information LOINCs offer with respect to the method applied in a laboratory test.
Supported standards for LabAnalyzer are identifiers from the Global Medical Device Nomenclature (GMDN), the European Medical Device Nomenclature (EMDN), and the Global Unique Device Identifier Database (GUDID). These identifiers specify the analyzer and test kit/reagent used during a laboratory analysis of interest. The Global Unique Device Identification Database (GUDID) specifies medical devices on the product level by a unique identifier, while the Global Medical Device Nomenclature (GMDN) or the European Medical Device Nomenclature (EMDN) provide type identifiers.
All new terminologies have been translated to RDF-format with the support of DCC and SIB.
Available resources
Contact persons:
Alexander Leichtle, Harald Witte
The Swiss BioRef ontology is available here:
- https://git.dcc.sib.swiss/sphn-semantic-framework/sphn-funded-projects/-/blob/main/SwissBioRef/SwissBioRef_ontology_2022-2__2022-05-20.ttl
- https://www.biomedit.ch/rdf/sphn-ontology/swissbioref
Information on terminologies used in SPHN and BioRef are provided by the SPHN Data coordination center:
For using the GMDN, please request a license from the GMDN Agency (https://www.gmdnagency.org/)
4. csv-RDF-converter, incl. QC-preprocessing module
Resource Description Framework (RDF) is the data format encouraged by SPHN projects due to the ability to integrate data from multiple sources effectively. Despite its benefits to facilitate data interoperability, RDF is not yet common across the board, in particular in smaller hospitals or institutions. We have therefore extended the general RDF-converter developed at Inselspital to accept csv-input for use on the central project space at BioMedIT. It is in principle portable to hospitals outside the SPHN-domain as well. The converter allowed Swiss Paraplegic Research and the University Children's Hospital Zurich to provide their data contribution not only to Swiss BioRef Central (the centralized approach), but also to Swiss BioRef TI4Health (the decentralized approach) which indirectly uses RDF-data via an i2b2- conversion step.
The csv-RDF-converter is a Python framework which enables the conversion of tabular data (csv-format) to subject-predicate-object triples of the RDF data model. It is based on the RDF-converter implemented at Inselspital which works with input from databases from the Clinical Data Warehouse. The tabular data must contain all elements required for the converter to produce meaningful output. Before conversion, a preprocessing step and quality check is run on the csv-data, ensuring for example valid ICD-10 codes and UCUM units.
The csv-RDF-converter uses the csv-data, an ontology, a corresponding mapping table, and a configuration file as input. The latter two define the concepts of interest for the conversion. The converter is able to process multiple concepts in parallel to speed up conversion on a high performance-cluster, and breaks down large data input into chunks. Turtle-files for each concept of interest are the output and match the input ontology. After conversion, the files are ready for RDF-i2b2-conversion and subsequent upload to a TI4Health instance.
Available resources
Contact persons:
Alexander Leichtle, Harald Witte
Github link to Inselspital-RDF-converter (Please request access from idsc@insel.ch): https://github.com/IDSC-io/sphn-rdf
For further information on the csv-RDF-add-on contact info@bioref.ch.
5. RDF-i2b2 ontology and data converter
Conversion to i2b2-format is a prerequisite to make interoperable RDF-data accessible to MedCo / TI4Health. A preexisting initial version of an RDF-i2b2-converter developed during the MedCo-project was able to process data in line with the 2020-release of the SPHN ontology. During Swiss BioRef, the RDF-i2b2 data converter has been enhanced at CHUV to be able to process RDF-data in line with the SPHN-ontology, release 2021-2.
The RDF-i2b2 converter is a Python framework allowing conversion from RDF knowledge graphs to CSV tables as intended for an i2b2 database system. It features two modules, an ontology converter and a data sample converter:
- Ontology converter: This converter turns an RDF metadata graph into a tree-structured list of absolute paths and identifying codes, representing a hierarchy the user can browse through using the Medco webclient. It takes as input the RDF knowledge graphs for the project ontology as well as the external terminologies (ICD-10, SNOMED, LOINC, etc.), also formatted in RDF. The converter extracts only the elements required for the project, using a couple of configuration files to filter and aggregate items if necessary. The ontology tables can be reused throughout all sites participating in a multicenter study.
- Data sample converter: This converter accepts RDF instances (consistent with the RDF metadata graph used as input to the ontology converter) and creates i2b2 observations from them. It assigns a unique identifying code to each observation, binding it to an existing ontology item. For Swiss BioRef, post-production routines were implemented, such as merging specific quantitative and qualitative observations to improve the user experience with respect to RDF.
A quality-check after the conversion allows to detect inconsistencies in the underlying data, for example, invalid or deprecated codes. After conversion, the tables can be incorporated in an i2b2 database and consequently loaded by MedCo/TI4Health.
Available resources
Contact persons:
Alexander Leichtle, Jules Fasquelle
Github link to RDF-i2b2 data converter (open-source):
Publication on RDF-i2b2 converter: https://pubmed.ncbi.nlm.nih.gov/35612099/
6. MedCo enhancements
MedCo was initially designed to process count data, e.g., from survival analysis. Both back-end and front-end of MedCo have been enhanced during Swiss BioRef by CHUV and Inselspital to process numerical data for statistical analysis and estimation of reference intervals. The modifications applied to MedCo include a new full-stack feature and several enhancements. The back-end of the said feature reuses the native selecting and counting of a population based on medical criteria. It adds to it the retrieval of laboratory result values for the selected population and the aggregation and processing of those results for statistical operations. This development required adding code accessing directly to the underlying database as well as API changes, in order to return multidimensional vectors instead of single counts. The front-end has been adapted accordingly and enhanced to help the user build study cases (including default filters, report export (PDF), or user group management). The results of the statistical queries are displayed as histogram including reference intervals and their confidence intervals. Search options in the ontology panel have also been enhanced to allow string-based term search. All modifications are available open-source and have been incorporated into the TI4Health codebase for use in future SPHN-projects and beyond.
Available resources
Contact persons:
Alexander Leichtle, Jean Louis Raisaro
Github-link to MedCo-variant tailored to Swiss BioRef (Swiss BioRef fork of MedCo; opensource):
https://github.com/CHUV-DS/glowing-bear-medco-bioref
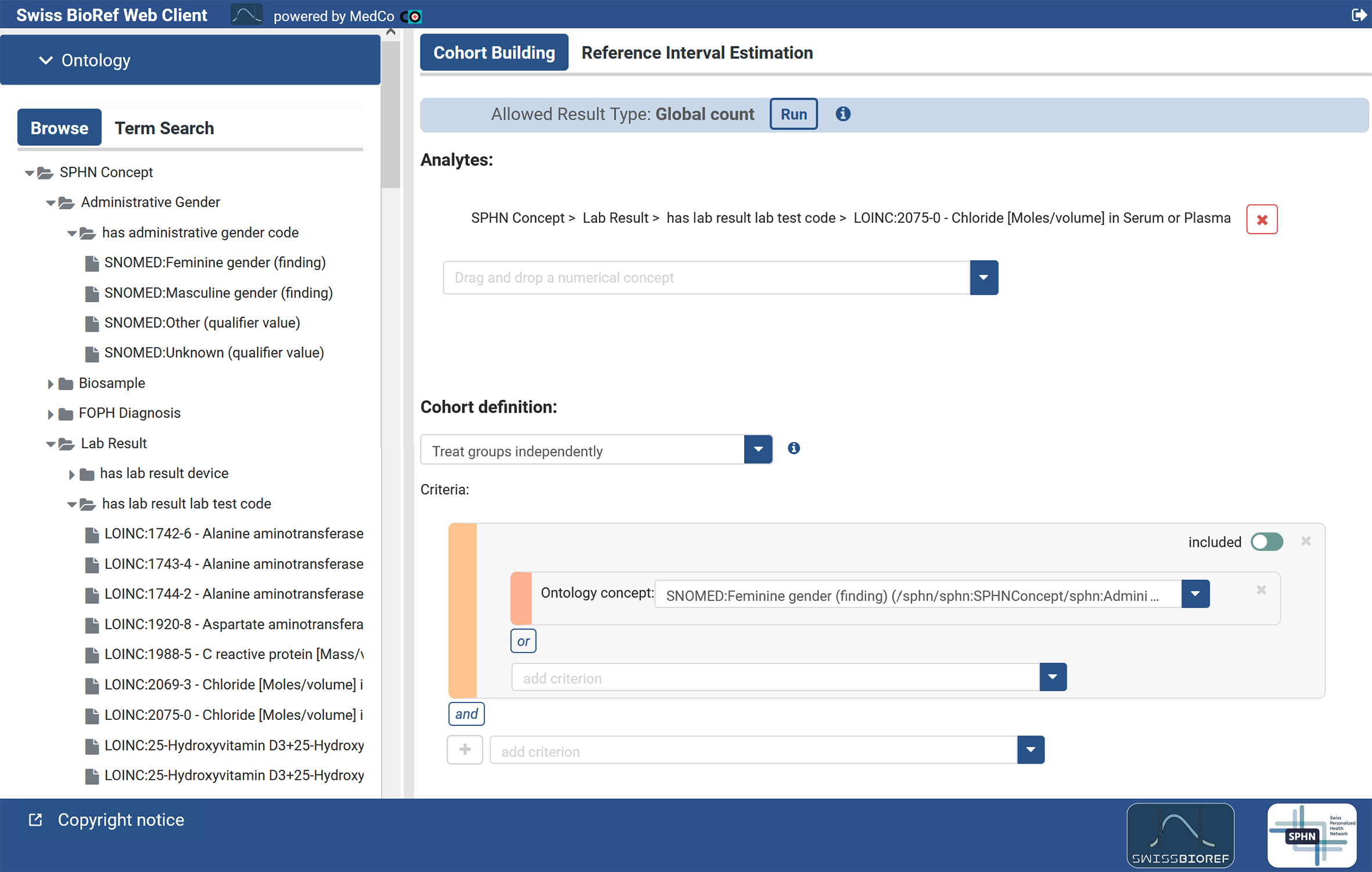
Figure 3: Swiss BioRef-MedCo user interface
Data dictionaries
For more information on the data dictionaries used within Swiss BioRef, please download the Table 2.
Follow up projects – continuation – next steps:
The ongoing BioRef-TI4Health pilot is a seamless continuation of Swiss BioRef. It aims at the industrialization of MedCo in the form of TI4Health, a MedCo-version developed further by the EPFL-spin-off Tune Insight. BioRef-TI4Health is a collaborative effort of Inselspital, CHUV, Tune Insight, DCC and BioMedIT.
The Swiss BioRef Consortium has been set up to persist after the completion of Swiss BioRef, hence the pilot can leverage the pre-established governance structures, including consortium agreement, DTUA and DTPA. Alike, the ontology developed during Swiss BioRef and the existing data set can be used.
Enhancements of MedCo (upgrade of RDF-i2b2 data converter to SPHN ontology release 2021-2; adaption of MedCo from processing simple count data (e.g. in survival analysis) to numerical data) have been integrated into the MedCo main repository from EPFL for open access and are currently being integrated into the TI4Health codebase by Tune Insight. The infrastructures established at BioMedIT during Swiss BioRef are used and extended during the pilot. For example, the preparations for MedCo-deployment on LeoMed at the Zürich BioMedIT node are directly usable for TI4Health-deployment. This includes security assessments, test deployments and establishment of reverse proxy connections. Moreover, secure access to dockerized applications run on LeoMed has been established, using reverse proxy and SWITCH edu-ID-based two-factor authentication (2FA).
During the Swiss BioRef project an RShiny app for the estimation of reference values from laboratory data with various distributions has been implemented. It supports the development of algorithms tailored to the distributed computing solutions (MedCo or TI4Health, respectively) and allows validation of results and performance. A docker container of the RShiny web application has been deployed on LeoMed and is accessible to Consortium members and registered users by 2FA-access.
It is envisioned to enable a broader audience to access the Swiss BioRef infrastructure, e.g., clinical specialists or researchers from the consortium members and possibly beyond. To ensure sustainability and long-term maintenance of the infrastructure, the project team is currently also reaching out to partners beyond academia for funding opportunities, including industry and clinical medicine societies.
Links and references relevant to the project
A manuscript describing the project setup and infrastructure architecture in detail has been submitted to BMJ Open and is currently under review. A preprint is available on medRxiv. The Swiss BioRef project website is currently being created (bioref.ch).
Presentations of the Swiss BioRef project:
- online presentation for a lecture series of the Swiss Institute of Bioinformatics (SIB) in collaboration with the SPHN (https://www.youtube.com/watch?v=R_j8MvM5A2A)
- several seminar series at the Inselspital and the University of Bern
- Nexus Personalized Health Technologies 2022 conference at ETHZ (poster presentation)
- 2nd Joint Personalized Health Day Switzerland in Bern (poster presentation)
- oral presentation as one of five selected talks at the Annual Assembly 2022 of the Schweizerische Gesellschaft für Klinische Chemie (SGKC).
Further information
- Task Force on Global Reference Interval Database (TF-GRID): https://www.ifcc.org/executive-board-and-council/eb-task-forces/task-force-on-global-reference-interval-database-tf-grid/
- Global Medical Device Nomenclature (GMDN): https://www.gmdnagency.org/
- Global Unique Device Identification Database (GUDID): https://accessgudid.nlm.nih.gov/
- European Medical Device Nomenclature (EMDN): https://webgate.ec.europa.eu/dyna2/emdn/
Watch the SPHN webinar
Presenter: Prof. Dr. med. Alexander Leichtle, Inselspital – Bern University Hospital
Disclaimer: The contents on this website are intended as a general source of information and have been provided by the project PIs. The SPHN Management Office is not responsible for its accuracy, validity, or completeness.

